Supel™ Swift HLB SPE Cartridges
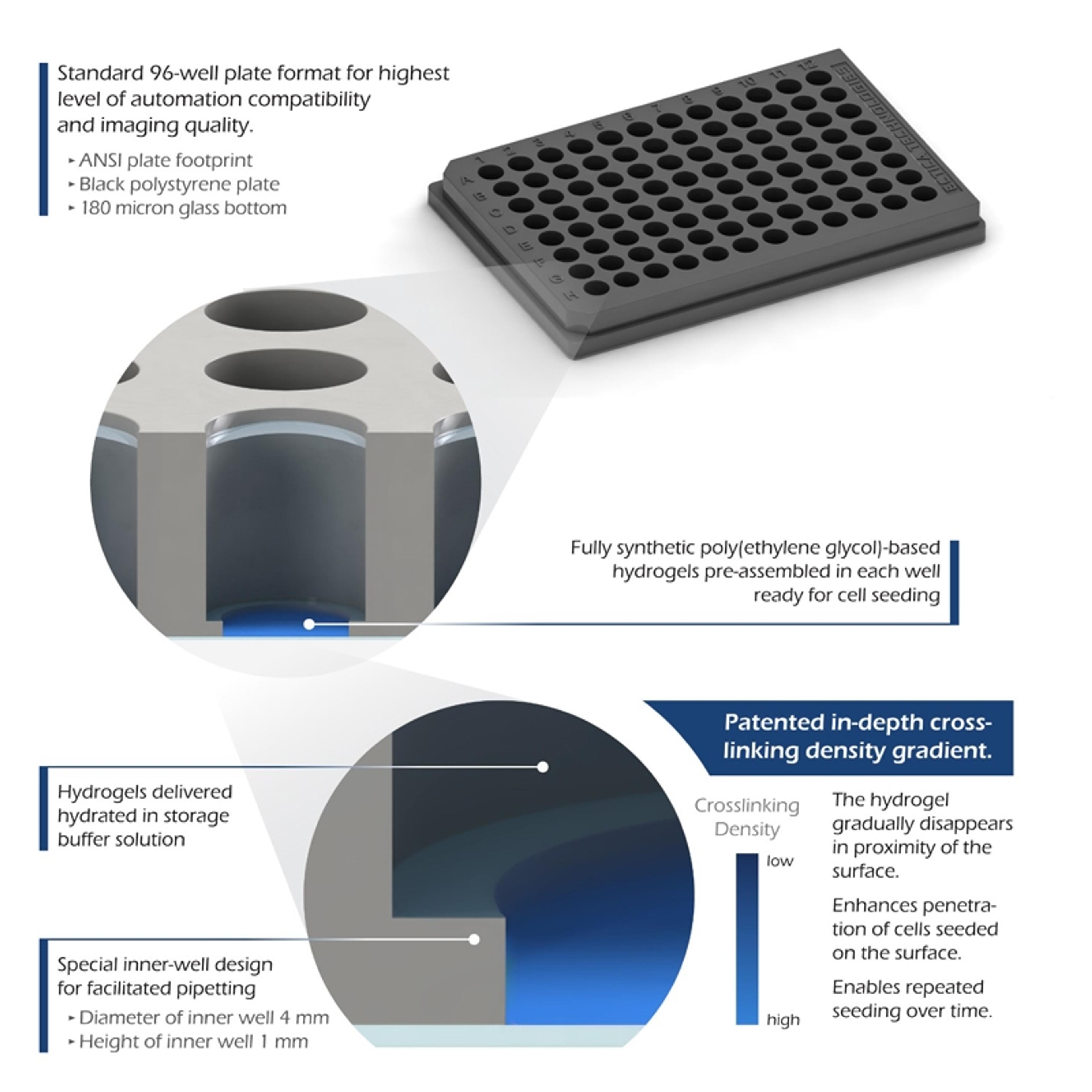
MerckSupel™ Swift HLB is a polymeric stationary phase for solid phase extraction (SPE) prior to instrumental analysis. It has both hydrophilic and lipophilic functional groups for the extraction of a broad range of compounds from aqueous samples. It retains analytes having diff erent polarities and Log P values due to its hydrophilic and lipophilic balance (HLB) property.